Evaluation and Benchmarking#
Five lessons that treat decoding evaluation as a core skill, drawing on MOABB (Chevallier, Aristimunha et al. 2024) and the evaluation guidance in Cisotto and Chicco (2024). Difficulty 2-3; assumes the core workflow track and the leakage-safe split lesson in particular.
Evaluation is where most EEG decoding claims fall apart: the model
trained on a single subject does not generalise to a held-out one, the
single-split accuracy hides session drift, and a paired comparison
between two pipelines is replaced by a bar chart with no statistics.
This category builds, in order, from a single within-subject split
toward a benchmark-grade paired comparison: which evaluation regime is
honest for your claim, and how do you report it. Sourced from
docs/tutorial_restructure_plan.md Category F (lines 425-442).
What you will learn:
When within-subject diagnostics are appropriate, and when they are marketing.
How to run a cross-subject evaluation – the gold standard for any generalisation claim – with
GroupKFoldand EEGDash’s split helpers.How to detect calibration drift across sessions of the same subject.
How to plot a learning curve as a function of training subjects, trials, or windows, and read it for sample-efficiency claims.
How to compare two pipelines on the same split and report the paired-Wilcoxon p-value the right way.
Run the lessons in order:
plot_50_within_subject_evaluation.py– single-subject diagnostics.plot_51_cross_subject_evaluation.py– the gold standard for generalisation.plot_52_cross_session_evaluation.py– calibration drift across sessions.plot_53_learning_curves.py– performance vs training data size.plot_54_compare_two_pipelines.py– paired comparison with the Wilcoxon test.
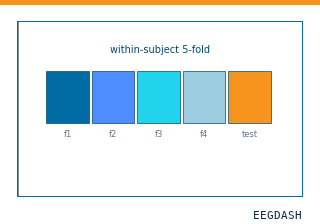

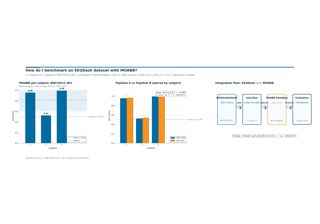
When is within-subject decoding the right scientific question?
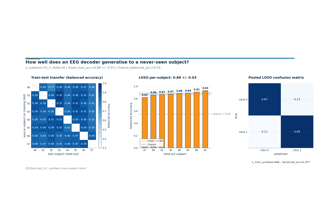
How well does an EEG decoder generalise to a never-seen subject?
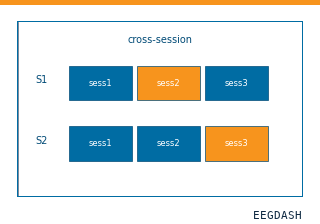

How much does a within-session decoder drift across sessions of the same subject?
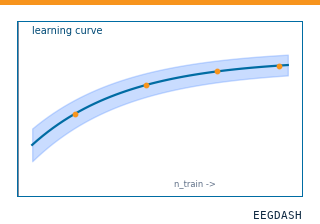
How does decoding accuracy scale with training-set size, and where does the curve plateau?
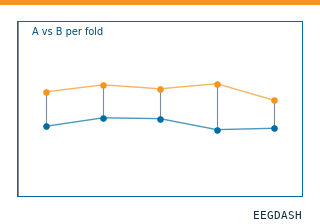
Is Pipeline A really better than Pipeline B, or did it luck out on one subject?