Feature Engineering#
EEGDash’s feature extraction package as a first-class option, not an afterthought to deep learning. Three lessons cover feature tables from windows, preprocessor / dependency trees that avoid recomputation, and a scikit-learn / LightGBM baseline straight from a feature table. Difficulty 1-2; assumes the core workflow track.
Hand-crafted features remain the strongest baseline on small EEG
datasets. They are interpretable, sample-efficient, and reproducible,
and a feature pipeline is the right tool the moment you have fewer than
a few thousand windows per class. The dependency-tree lesson teaches
why you should compute the spectrum once and derive band powers from
it, rather than recomputing the FFT five times. Sourced from
docs/tutorial_restructure_plan.md Category E (lines 442-458).
What you will learn:
How to extract a small feature table (band power, statistical moments, Hjorth parameters) from windows of an
EEGDashDataset.How to compose preprocessors and feature definitions as a tree where shared computations (FFT, covariance) execute once.
How to feed the feature table into
sklearn.pipeline.Pipelineor LightGBM and run a stratified cross-validation.How to compare a feature baseline against a deep model on the same split (the Evaluation track builds on this).
Run the lessons in order:
plot_40_first_features.py– a feature table from windows.plot_41_feature_trees.py– preprocessor and dependency trees.plot_42_features_to_sklearn.py– a scikit-learn / LightGBM baseline from feature tables.
How do I turn EEG windows into a band-power feature matrix?
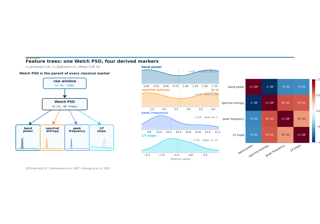
How do classical EEG markers compose on top of one Welch PSD?

How do I push EEGDash features through a scikit-learn Pipeline?